LD Block-Based Genotype Embeddings for Asthma Biology and UK Biobank Extension (Ongoing)
Built a two-stage LD-block representation learning framework for asthma genotype data, combining block-wise VAEs with transformer-based contextualization to learn biologically meaningful subject embeddings.
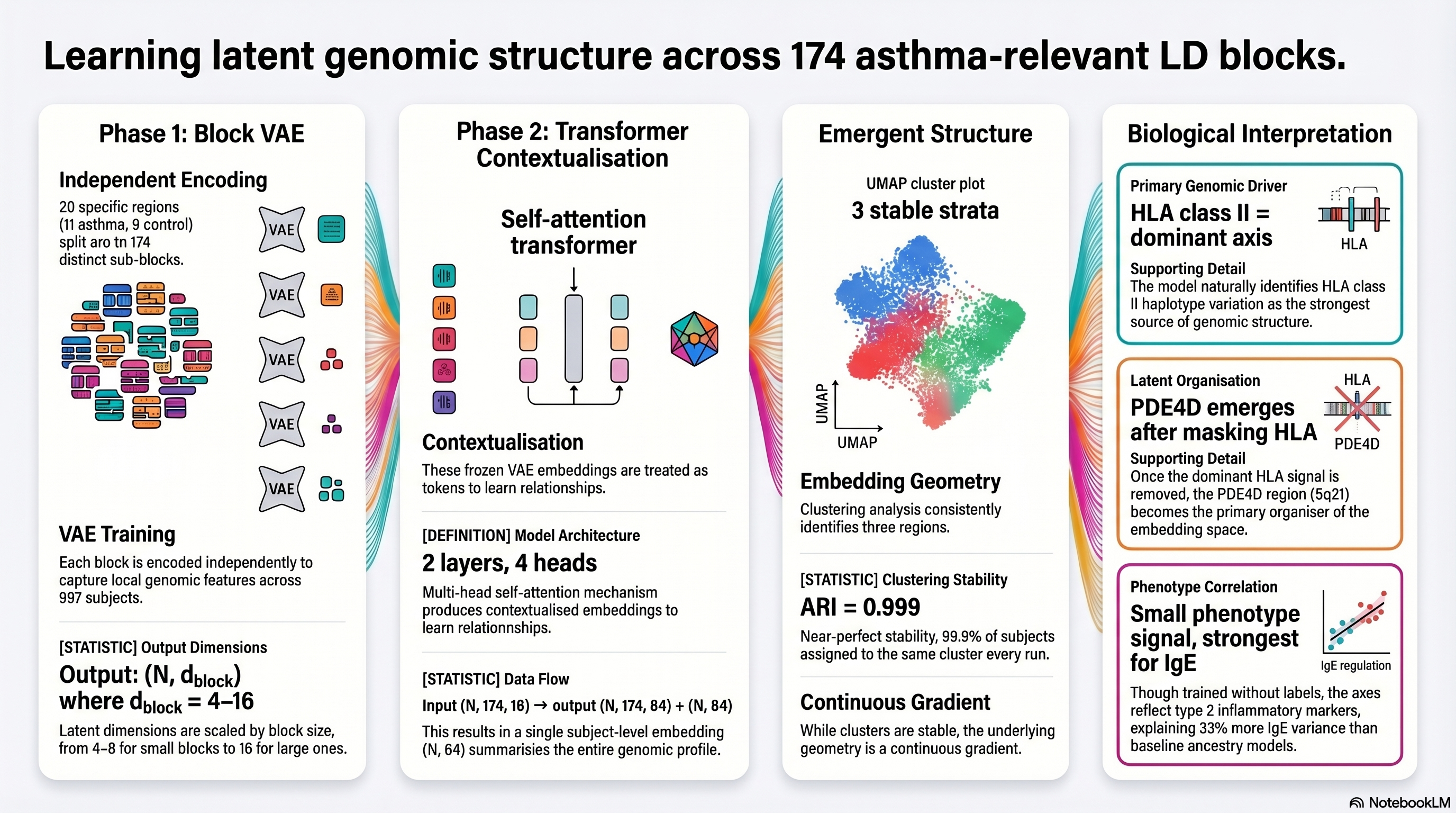
Two-stage LD block embedding framework showing independent block encoding, transformer contextualization, emergent genomic structure, and biological interpretation including HLA class II dominance and PDE4D emergence.
Key insights
- Recovered known structure: HLA class II emerged as the dominant unsupervised axis, showing that the model recovers meaningful asthma biology without using phenotype labels. Found secondary biology: After masking HLA effects, PDE4D appeared as an independent signal, suggesting the framework can reveal additional disease-relevant pathways beyond the strongest locus.
- Transformer adds value: Cross-block contextualization preserved local LD structure while reorganizing subjects into clearer biological patterns than block-level embeddings alone.
- Interpretable attribution: Used leave-one-block-out perturbation analysis to quantify how each LD block contributed to the learned subject embeddings and latent genomic structure.
- Scaling next: The same framework is now being extended to UK Biobank cohort.
Single-Cell Immune State Modeling and Multi-Modal Integration
Built single-cell analysis workflows spanning RNA-only PBMC benchmarking and ongoing paired RNA+ATAC integration. Combined clustering, cell-state scoring, cell–cell communication, chromatin accessibility, and transcription-factor inference to characterize immune-cell states across stimulated and control conditions.
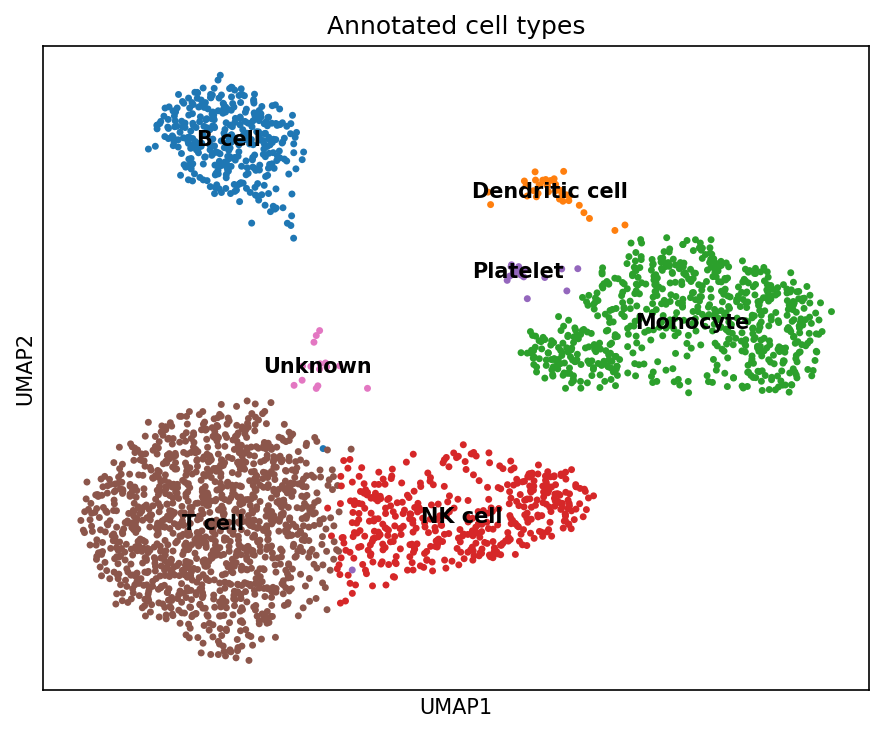
UMAP of PBMC single-cell RNA-seq data after clustering and annotation, highlighting major immune-cell populations used for downstream cell-state scoring, differential expression, and multi-modal integration analyses.
Highlights
- RNA-only benchmarking: Compared multiple cell-state scoring strategies across PBMC3k and interferon-stimulated Kang datasets using clustering, marker genes, and immune program scores.
- Cross-modal integration: Integrated paired RNA+ATAC data with weighted nearest neighbors (WNN) to evaluate how chromatin accessibility refines immune-cell separation.
- Regulatory modeling: Inferred transcription-factor activity from RNA and validated it against ATAC motif accessibility, with ongoing work on pseudotime and dynamic regulatory programs.
Proteomics Analysis of Long COVID Heterogeneity
Built a proteomics analysis workflow to study neurocognitive Long COVID using Olink NPX data, combining covariate-adjusted modeling in a pediatric cohort with scalable phenotype–proteomics integration in ~50,000 UK Biobank participants.
Highlights
- Subtype-focused analysis: Distinguished neurocognitive and non-neurocognitive Long COVID using protein-level signatures.
- Replication at scale: Extended the workflow to ~50K UK Biobank participants with distributed Spark-based integration.
- Biological follow-up: Candidate proteins and pathway-level summaries support immune and neuroinflammatory hypotheses for Long COVID heterogeneity.
Endophenotype-driven GWAS Framework for Asthma Heterogeneity
Developed a subtype-aware GWAS framework for asthma by combining PCA-derived clinical endophenotypes with categorical association modeling, enabling genetic analysis across biologically coherent patient subgroups rather than treating asthma as a single uniform phenotype.
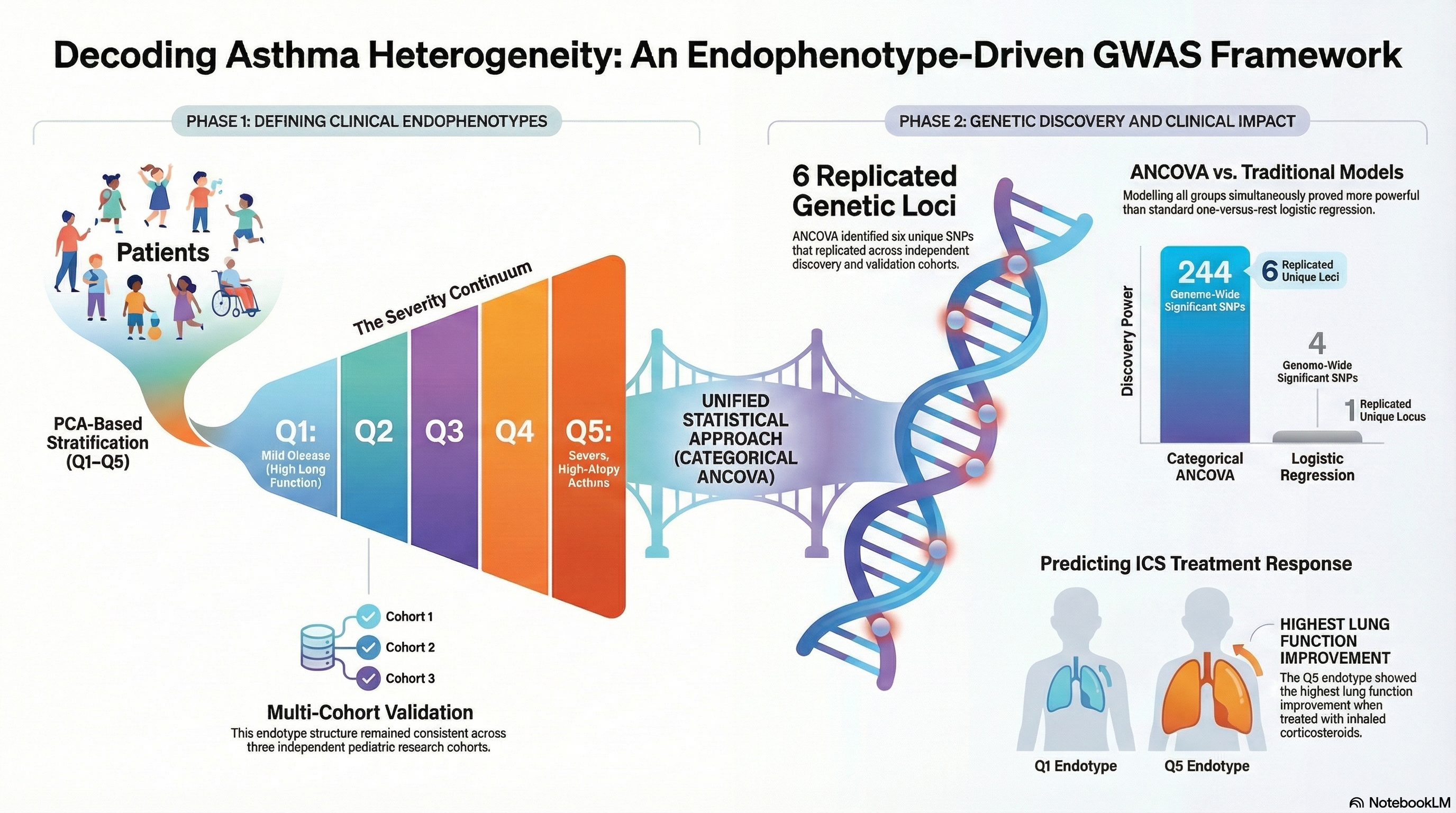
PCA-based clinical stratification into endophenotypes integrated with subtype-aware GWAS to identify asthma-associated loci across disease severity groups.
Highlights
- Subtype-aware association testing: Endophenotype-based modeling improved detection of genetic signals compared with standard case-control GWAS.
- Replication across cohorts: Multiple loci, including DGKI and MIR99AHG, replicated in independent asthma datasets.
- Clinical relevance: Endophenotypes aligned with severity, IgE, lung function, and differential ICS treatment response.
miRNA Modifiers of Inhaled Corticosteroid Response in Asthma
Identified miR-584-5p as a potential biomarker of inhaled corticosteroid resistance in asthma using interaction modeling across two pediatric cohorts.
Machine Learning Analysis of Pediatric COVID-19 Clinical Data
Built a Random Forest model integrating chest X-ray impressions, symptoms, and demographics to predict pediatric COVID-19 infection status.
Structured Modeling of Context-Aware Systems
PhD research on modeling event-driven mobile applications using probabilistic sequence models, neural representation learning, and combinatorial optimization.