DNA Sequencing Explained: What Does Reading the Genome Mean?
Reading Path: Step 1 — Measurement Foundations (Start Here)
This article is part of the Foundations of Genomic Data series.
What does DNA sequencing capture?
DNA sequencing refers to determining the order of nucleotides along a DNA molecule. In practical terms, it produces a digital record of the sequence present in a biological sample.
Sequencing does not target genes specifically. It reads whatever DNA fragments are present, including protein-coding regions, noncoding regions, regulatory elements, and repetitive sequences.
For this reason, DNA sequencing describes potential sources of biological variation rather than biological activity itself.
Most of the genome is not protein-coding
It is often assumed that sequencing is mainly about genes. In reality, protein-coding exons make up only a small portion of the human genome.
The figure below summarizes the approximate composition of the genome. A large fraction consists of intronic, regulatory, and repetitive sequences, while protein-coding regions represent a comparatively small share.
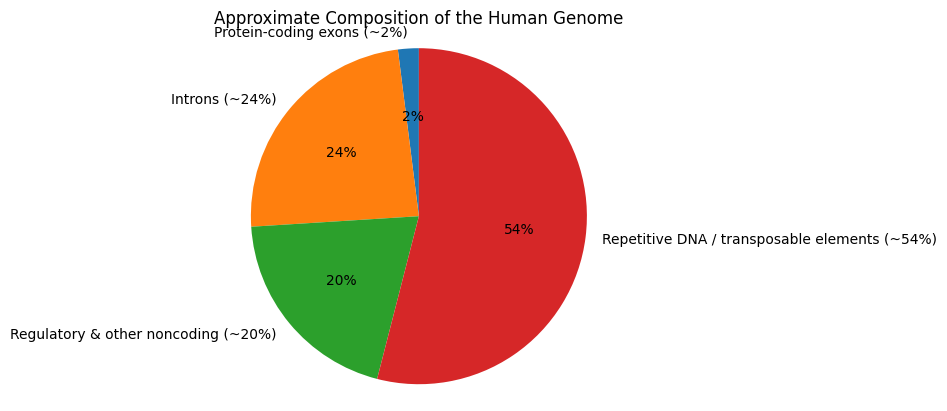
This matters because most variants identified through sequencing lie outside protein-coding regions. Their effects, when present, are often mediated through gene regulation, chromatin organization, or broader genomic context.
Types of DNA sequencing
DNA sequencing can be applied at different scales, depending on the scientific or clinical question.
Whole Genome Sequencing reads most of the genome and provides the broadest view of genetic variation, including noncoding and structural variants.
Whole Exome Sequencing focuses on protein-coding regions. While narrower in scope, it is commonly used in settings where coding mutations are of primary interest.
Targeted sequencing restricts analysis to predefined genomic regions. This approach is typically chosen when prior knowledge suggests specific loci are relevant.
Each approach reflects a balance between coverage, cost, and interpretability rather than a hierarchy of completeness.
What DNA sequencing is well suited for
DNA sequencing is particularly useful for identifying inherited genetic variation. Common applications include:
Detection of single-nucleotide variants and small insertions or deletions.
Identification of larger structural variants.
Analysis of rare genetic mutations.
Population-scale studies of genetic diversity.
Because DNA sequence is largely stable across cells and over time, sequencing provides a consistent reference layer for downstream analyses.
What DNA sequencing does not measure
DNA sequencing does not indicate whether a gene is active, repressed, or dynamically regulated. Individuals with similar DNA sequences can exhibit very different cellular behavior.
This limitation reflects the biological role of DNA. Sequencing captures the genetic substrate, not how it is used in a particular cell or condition.
Why sequencing is only one layer of genomic data
DNA can be thought of as a stable blueprint. Many downstream processes determine which parts of that blueprint are read, when, and in which cells. RNA sequencing is designed to measure this active layer of gene regulation. For this reason, it is often combined with RNA sequencing, epigenomic assays, and other measurements.
Later articles in this series explore how these complementary data types address different biological questions and how they are integrated in practice.
Continue Reading →