From DNA to Disease: A Layered View of Biology
Biology is often taught in separate pieces due to its complexity: genetics, RNA, proteins, pathways, disease. In practice, these are not separate systems. They are layers of the same system.
A biologist rarely asks only, “Which gene matters?” The more useful question is:
- What changed?
- At which layer did it change?
- How does that change propagate through the system?
- Why does it lead to disease in one person but not another?
Biology behaves less like a linear chain and more like a large control system with sensors, feedback loops, amplifiers, and fail-safes. Genes, RNAs, proteins, and signals continuously influence one another, and disease emerges when enough of those interacting controls shift out of balance.
The framework I find most useful is:
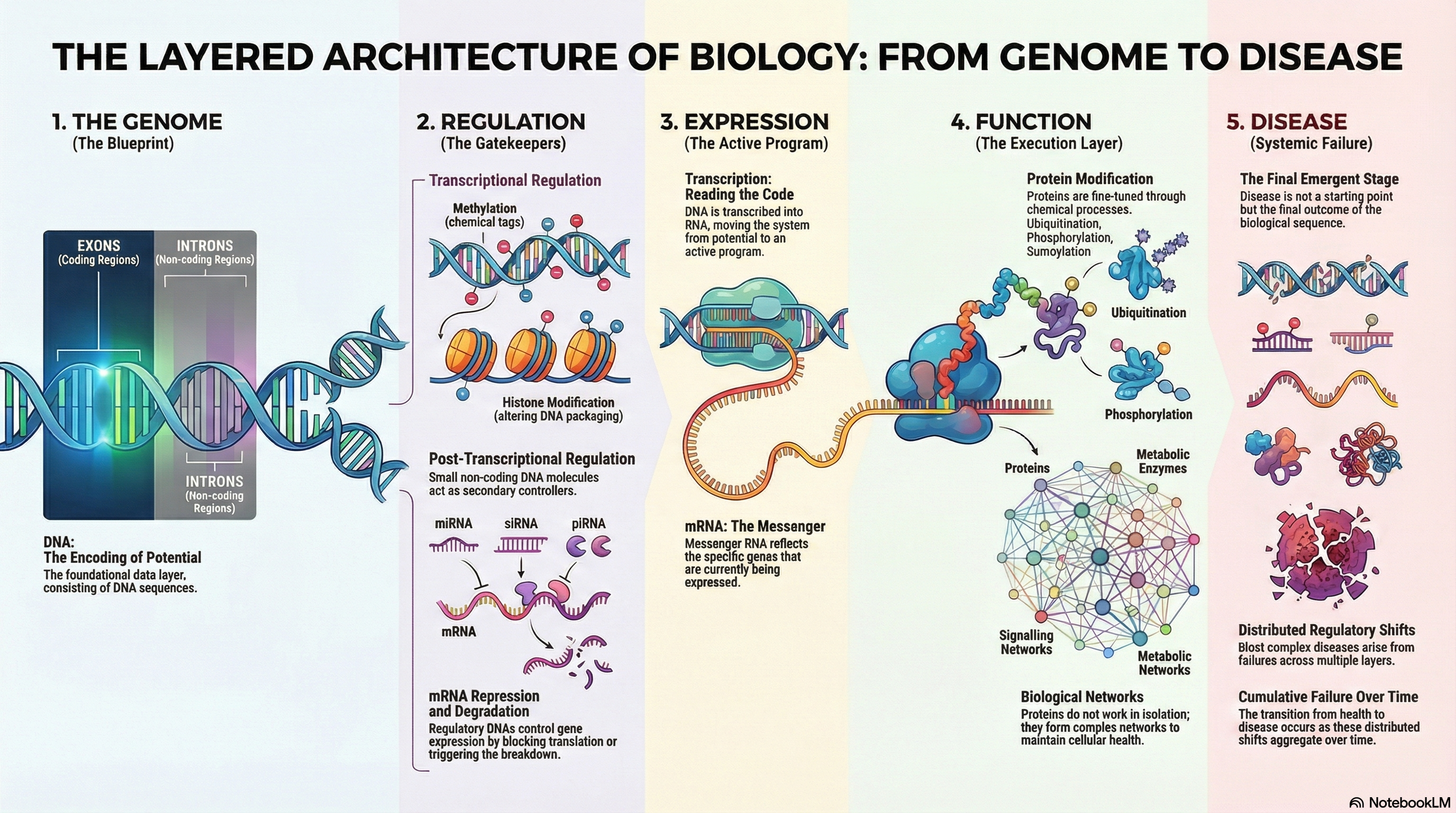
Genome → Regulation → Expression → Function → Disease
The genome provides potential. Regulatory systems determine which parts of that potential are accessible. RNA reflects what is active right now. Proteins carry out the work. Disease appears when enough small changes accumulate across these layers.
The Big Picture
Most discussions stop at the classic central dogma:
DNA → RNA → Protein
That is useful, but incomplete.
Biology is not just a one-way chain. DNA can be regulated before it is read. RNA can be controlled after it is made. Proteins can be modified after they are produced. Those changes interact through signaling and metabolic networks, and disease emerges from the combined effect.
A simple way to think about the layers:
| Layer | Main question |
|---|---|
| Genome | What could happen? |
| Regulation | What is allowed to happen? |
| Expression | What is happening right now? |
| Function | What is the cell doing? |
| Disease | What happens when the system shifts over time? |
No single dataset captures the whole story. DNA, RNA, and proteins each measure different parts of the same system.
1. The Genome: The Blueprint of Potential
The genome is the long-term information layer. DNA contains the instructions a cell could use, but not every instruction is active at the same time.
Some regions of DNA are coding regions, called exons. These are the parts that can eventually become proteins. Other regions are non-coding, including introns and regulatory DNA. Most of the genome is non-coding, but that does not mean it is unimportant. Many disease-associated variants sit in regulatory regions rather than in genes themselves.
DNA also has to be copied. During replication, the genome is duplicated so that cells can divide and pass the same information forward.
The key idea is that the genome represents potential, not certainty. Two people may carry the same variant, but only one develops disease because the downstream layers differ.
What is commonly analyzed at this layer?
| Common analyses | What they identify |
|---|---|
| Variant calling | Which DNA variants are present |
| GWAS | Which variants are associated with disease |
| Fine-mapping | Which variants are most likely causal |
| Polygenic risk scores | Overall inherited risk from many variants |
For example, a genome-wide association study may identify variants near asthma-related genes, but it usually does not explain how those variants actually change biology.
2. Beyond the Classic Central Dogma
The standard central dogma describes a one-way flow:
DNA → RNA → Protein
That is still broadly true, but the full system is more complex.
DNA can copy itself through replication. RNA can sometimes flow back into DNA through reverse transcription, which occurs in retroviruses and some mobile genetic elements.
More importantly, the layers influence one another. DNA affects RNA, but RNA and proteins also feed back into regulation. Biology behaves more like an interacting system than a straight line.
Regulation exists at every level:
- DNA can be chemically modified
- RNA can be repressed or degraded
- Proteins can be activated, tagged, or destroyed
This is why two cells with the same genome can behave very differently.
3. Regulation: The Gatekeepers
Genes are not simply “on” or “off.” Cells constantly decide which parts of the genome are accessible and when.
Regulation is the layer that controls that decision.
DNA-Level Regulation: Epigenetics
Before a gene can be used, the cell must be able to access it.
Two major mechanisms control this:
- DNA methylation
- Histone modification
DNA methylation adds chemical tags to DNA. Histone modification changes how tightly DNA is packaged around proteins called histones.
When DNA is tightly packed, genes are harder to read. When it is open, genes are easier to activate.
This is the basis of epigenetics: changes in gene activity without changing the DNA sequence itself.
RNA-Level Regulation
Even after RNA is produced, the cell can still control it.
Small non-coding RNAs such as miRNA, siRNA, and piRNA can bind to messenger RNA and either block it or trigger its degradation.
As a result, a gene may be transcribed, but its message never becomes protein.
This is one reason why measuring RNA alone does not always tell the full story.
Protein-Level Regulation
Proteins are also regulated after they are made.
Common protein modifications include:
- Phosphorylation
- Ubiquitination
- Sumoylation
These modifications can activate a protein, change where it moves, or mark it for destruction.
A protein may be present, but inactive. Another may be short-lived because it is rapidly degraded. The amount of protein is only part of the story.
What is commonly analyzed at this layer?
| Common analyses | What they identify |
|---|---|
| DNA methylation analysis | Which regions of DNA are more or less accessible |
| ChIP-seq / histone profiling | Which genes are likely to be active or repressed |
| ATAC-seq | Which regions of the genome are open |
| miRNA analysis | Which RNAs are suppressing gene activity |
| eQTL analysis | Which genetic variants affect gene expression |
These analyses help answer a different question from genetics alone: not just which variant exists, but how it changes the behavior of the cell.
4. Expression: The Active Program
DNA contains all possible instructions. RNA reflects the instructions currently being used.
When a gene is active, the cell transcribes DNA into messenger RNA (mRNA). That RNA acts as the working copy of the gene.
Because of this, RNA gives a snapshot of what the cell is doing at a particular moment.
The same genome can produce very different expression patterns in different tissues, conditions, or diseases. A lung cell and a neuron have the same DNA, but they express very different genes.
In disease, expression patterns often shift. Some genes become more active. Others become less active. Looking at those shifts can help identify the pathways involved.
What is commonly analyzed at this layer?
| Common analyses | What they identify |
|---|---|
| RNA-seq differential expression | Which genes change between groups |
| PCA or clustering | Whether samples form distinct subgroups |
| Pathway enrichment | Which biological processes are more active |
| Single-cell RNA-seq | Which cell types are driving the signal |
For example, two patients may both have asthma, but their expression profiles may show that different pathways are active. One may have a stronger inflammatory program, while another may have a stronger epithelial response.
5. Function: Proteins and Networks
RNA reflects the program, but proteins carry out the work.
Proteins build structures, transmit signals, and perform metabolic reactions. However, they rarely act alone. Most proteins work as part of signaling pathways, metabolic pathways, and larger interaction networks.
Because of this, a small change in one gene can spread through the system.
A variant may slightly change the expression of one protein. That protein may affect a signaling pathway. The pathway may alter many downstream genes. Eventually, the combined effect may appear as disease.
This is why common diseases rarely result from one broken molecule. More often, they arise from a network that has shifted out of balance.
What is commonly analyzed at this layer?
| Common analyses | What they identify |
|---|---|
| Proteomics | Which proteins are increased or decreased |
| Phosphoproteomics | Which proteins are active or modified |
| Protein-protein interaction networks | Which proteins work together |
| Pathway analysis | Which signaling or metabolic systems are altered |
This layer is often where separate molecular changes begin to connect into a single biological explanation.
6. Disease as an Emergent Property
Disease is not usually the direct result of one event.
Instead, it emerges when many small changes accumulate across layers:
- Genetic variants
- Epigenetic changes
- RNA regulation
- Protein activity
- Network rewiring
The final phenotype is the visible result of those combined shifts.
This is especially true for common diseases such as asthma, diabetes, cancer, and many neurological disorders. In these diseases, there is rarely one “bad gene.” Instead, there are many small changes that together move the system toward disease.
A useful way to think about it is:
- The genome sets the possibilities
- Regulation determines which possibilities are used
- Expression shows what is active
- Proteins carry out the response
- Disease appears when the balance breaks down
Disease also develops over time. The system may compensate for small changes for years before symptoms appear. By the time disease becomes visible, the underlying biology may have been shifting for a long time.
Why Measuring One Layer Is Not Enough
Different technologies measure different parts of the system.
| Layer | Typical measurements |
|---|---|
| Genome | DNA sequencing, genotyping arrays |
| Regulation | Methylation assays, ATAC-seq, ChIP-seq, miRNA profiling |
| Expression | RNA-seq |
| Function | Proteomics, phosphoproteomics |
| Disease | Clinical traits, imaging, symptoms |
Each measurement is useful, but incomplete on its own.
A genetic study may identify a disease-associated variant. An RNA study may identify a changed pathway. A proteomics study may show which proteins are active. The strongest explanation usually comes from placing those layers together.
This is the idea behind multi-omic integration: connecting DNA, RNA, proteins, and phenotype into a single model of disease.
Where the Story Gets More Complex
The simple Genome → Regulation → Expression → Function → Disease framework is useful, but real biology contains additional layers.
Epigenomics asks how chemical changes alter gene accessibility. Three-dimensional genome organization asks how DNA folding changes which genes can interact. Multi-omic integration asks how all of the layers fit together.
Most importantly, disease is not static. These layers interact over time.
Understanding disease requires more than measuring one layer. It requires understanding how the layers influence one another.